Setup a Private Local RAG Knowledge Base

To build a private Retrieval-Augmented Generation (RAG) system, pair a local vector database like Qdrant with an embedding model like BGE-M3 . Add a local LLM through Ollama , and you can index hundreds of documents and ask questions about them. Your data stays on your machine.
Why RAG? The Problem With Pure LLM Memory
Large language models sound smart, but they are poor knowledge stores. They learn from old training data and know nothing about files you created later or keep private. Ask about your own data, and the model will often guess. Even strong open weight models like Llama 4.0 can invent plausible but wrong answers about content they never saw. For a deeper breakdown of why LLM hallucinations happen and how to measure them, the issue goes beyond missing context.
The simple workaround is to paste all your documents into one context window. That fails fast. Context windows are limited, long prompts are slow and costly, and models often miss material in the middle. Research calls this the “lost in the middle” problem. If you stuff a 200-page legal document into one prompt, the model may skip the clause on page 95.
RAG avoids those problems by changing how retrieval works. First, you turn each document chunk into a vector and store it in a vector database. When a query arrives, the same model turns the query into a vector. The database then returns the 3-10 closest chunks. You place only those chunks in the LLM context. The model answers from retrieved evidence instead of memory. That makes the system more accurate, easier to trace, and easy to update without retraining. Once this base works, you can move to setups where the model picks its own sources and judges whether the results are enough.
RAG and fine-tuning solve different problems. Use RAG for knowledge that changes often, such as document sets, Obsidian vaults, codebases, and support archives. Use fine-tuning for fixed behavior, house style, or strict output shapes. If you need answers about your documents, RAG is almost always the right pick.
How To Build the Private RAG Pipeline
The build has seven steps. The outline below gives the short version. The rest of this post explains why each step helps and shows the code.
Build a Private Local RAG Knowledge Base
Install the Python dependencies
pip install qdrant-client fastembed ollama langchain-text-splitters pymupdf python-docx to pull the client, embedder, splitter, and document loaders into one venv.Pull a local LLM with Ollama
ollama pull mistral-nemo (or llama4 on a 24 GB GPU) so the generation step can answer entirely on your hardware.Start Qdrant in Docker with telemetry off
QDRANT__TELEMETRY_DISABLED=true and a mounted volume so the index survives restarts.Load BGE-M3 and create the collection
fastembed to load BAAI/bge-m3, then create a Qdrant collection of 1024-dimensional vectors with cosine distance for dense retrieval.Chunk and ingest your documents
RecursiveCharacterTextSplitter (512 tokens, 64 overlap), embed each chunk, and upsert into Qdrant with source filename and page payload.Add hybrid search and a reranker
Generate grounded answers with Ollama
Choosing Your Stack: Vector DB, Embedding Model, and LLM
Your RAG stack has three parts: the vector database, the embedding model, and the LLM. The parts affect one another. A poor mix can hurt retrieval quality or exceed your RAM.
For the database, Qdrant is the best pick for a serious local setup. It runs as one Docker image, saves data to disk, and supports dense plus hybrid sparse+dense search. That hybrid support helps retrieval quality. Chroma is easier for quick tests because it runs in Python and skips Docker, but it lacks stronger features such as filtering-aware HNSW indexing and built-in hybrid search. Milvus fits teams that index millions of documents. For a personal knowledge base, that is usually more than you need. The table below summarizes the tradeoffs:
| Feature | Qdrant | Chroma | Milvus | LanceDB |
|---|---|---|---|---|
| Deployment | Docker / binary | Python in-process | Docker / K8s | Python in-process / S3 |
| Hybrid search | Built-in (RRF) | No | Yes (BM25 plugin) | No |
| Disk-backed storage | Yes (HNSW on disk) | Yes (SQLite) | Yes | Yes (Lance columnar) |
| Filtering on metadata | Yes (payload filters) | Yes (basic) | Yes | Yes |
| Ease of local setup | Medium (Docker) | Very easy | Complex | Very easy |
| Best for | Production local RAG | Prototyping | Enterprise scale | Analytics workloads |
For embeddings, BGE-M3 from BAAI is a strong local default. It makes 1024-dimensional dense vectors, supports more than 100 languages, and can also produce sparse BM25 style vectors. That makes it a natural match for Qdrant hybrid search. all-MiniLM-L6-v2 is faster and smaller at 384 dimensions, so it works for quick prototypes. nomic-embed-text is a solid middle option with strong English quality at 768 dimensions.
| Model | Dimensions | Languages | Speed (CPU) | Quality |
|---|---|---|---|---|
| BGE-M3 | 1024 | 100+ | ~80ms/chunk | Excellent |
| nomic-embed-text | 768 | English-primary | ~45ms/chunk | Very good |
| all-MiniLM-L6-v2 | 384 | English | ~15ms/chunk | Good |
For generation, Ollama keeps local inference simple. Any model you can pull via ollama pull, including Llama 4.0, Mistral Nemo, or Gemma 3, can answer the final prompt. On a 16GB RAM system with no discrete GPU, Mistral Nemo (12B, Q4 quantized) is a good fit. On a machine with an RTX 50 series GPU and 24GB+ VRAM, Llama 4.0 at Q4 works well. Install the dependencies before you start:
pip install qdrant-client fastembed ollama langchain-text-splitters pymupdf python-docxPull the generative model via Ollama:
ollama pull mistral-nemoStart Qdrant with Docker:
docker run -d --name qdrant \
-p 6333:6333 -p 6334:6334 \
-v $(pwd)/qdrant_storage:/qdrant/storage \
qdrant/qdrant
Document Ingestion and Chunking Strategies
The quality of a RAG system depends heavily on chunking. Retrieval compares query vectors with chunk vectors. If one chunk mixes two topics because you split every 512 characters, its embedding becomes a muddy average. Then it may not rank well for either topic.
Fixed-size character chunking is fast and poor. It often cuts a sentence, table, or code block in half. That makes chunks hard to read and gives you weak embeddings. Even though many tutorials start here, avoid it in production.
Recursive character splitting is a solid default. LangChain’s RecursiveCharacterTextSplitter tries paragraph breaks first, then sentences, then words, and falls back to raw character counts only at the end. That keeps most chunks coherent. Start with 512 tokens and 64 tokens of overlap:
from langchain_text_splitters import RecursiveCharacterTextSplitter
splitter = RecursiveCharacterTextSplitter(
chunk_size=512,
chunk_overlap=64,
separators=["\n\n", "\n", ". ", " ", ""],
)
chunks = splitter.split_text(document_text)For code files (.py, .ts, .go), use language-aware splitting. LangChain provides Language.PYTHON, Language.JS, and similar options. They split on function and class boundaries instead of raw character counts, which keeps signatures with their bodies.
Different file types need different loaders. PDFs are the most common and the hardest. pymupdf (imported as fitz) keeps layout fairly well, handles many multi-column files, and extracts page numbers for citations. DOCX files work well with python-docx. Markdown from Obsidian vaults benefits from a splitter that respects heading structure.
Metadata is easy to skip and worth the effort. Each chunk in Qdrant should store the source filename, page or section number, heading, and a created or modified timestamp. That data lets you filter searches, such as “only search documents tagged as ’legal’,” and lets the system cite sources instead of returning plain text.
Here is a complete ingestion pipeline for a directory of PDFs:
import fitz # pymupdf
import os
from pathlib import Path
from langchain_text_splitters import RecursiveCharacterTextSplitter
from fastembed import TextEmbedding
from qdrant_client import QdrantClient
from qdrant_client.models import Distance, VectorParams, PointStruct
import uuid
COLLECTION_NAME = "knowledge_base"
EMBED_MODEL = "BAAI/bge-m3"
QDRANT_URL = "http://localhost:6333"
client = QdrantClient(url=QDRANT_URL)
embedder = TextEmbedding(model_name=EMBED_MODEL)
# Create collection if it doesn't exist
if not client.collection_exists(COLLECTION_NAME):
client.create_collection(
collection_name=COLLECTION_NAME,
vectors_config=VectorParams(size=1024, distance=Distance.COSINE),
)
splitter = RecursiveCharacterTextSplitter(chunk_size=512, chunk_overlap=64)
def ingest_pdf(pdf_path: Path):
doc = fitz.open(pdf_path)
all_points = []
for page_num, page in enumerate(doc, start=1):
page_text = page.get_text("text")
if not page_text.strip():
continue
chunks = splitter.split_text(page_text)
for chunk in chunks:
if len(chunk.strip()) < 50: # skip tiny fragments
continue
embedding = list(embedder.embed([chunk]))[0].tolist()
point = PointStruct(
id=str(uuid.uuid4()),
vector=embedding,
payload={
"text": chunk,
"source": pdf_path.name,
"page": page_num,
"path": str(pdf_path),
},
)
all_points.append(point)
# Upsert in batches of 100
for i in range(0, len(all_points), 100):
client.upsert(
collection_name=COLLECTION_NAME,
points=all_points[i : i + 100],
)
print(f"Ingested {len(all_points)} chunks from {pdf_path.name}")
docs_dir = Path("./documents")
for pdf_file in docs_dir.glob("**/*.pdf"):
ingest_pdf(pdf_file)Beyond Embedding: Semantic vs. Hybrid Search
Pure dense vector search has clear failure modes in production. Dense embeddings are good at concept matching, so they help with questions like “what does the contract say about liability?” They are weak at exact terms. Ask for “Section 4.2.1,” “the BGE-M3 model,” or “E_CONN_TIMEOUT,” and the right document may rank low because embeddings capture meaning, not exact strings.
BM25 sparse retrieval is the classic exact match method. It scores documents from term frequency and inverse document frequency. That makes it good for proper nouns, version numbers, model names, and error codes. It is much weaker at synonym and concept matching. On its own, it is not enough for a full knowledge base.
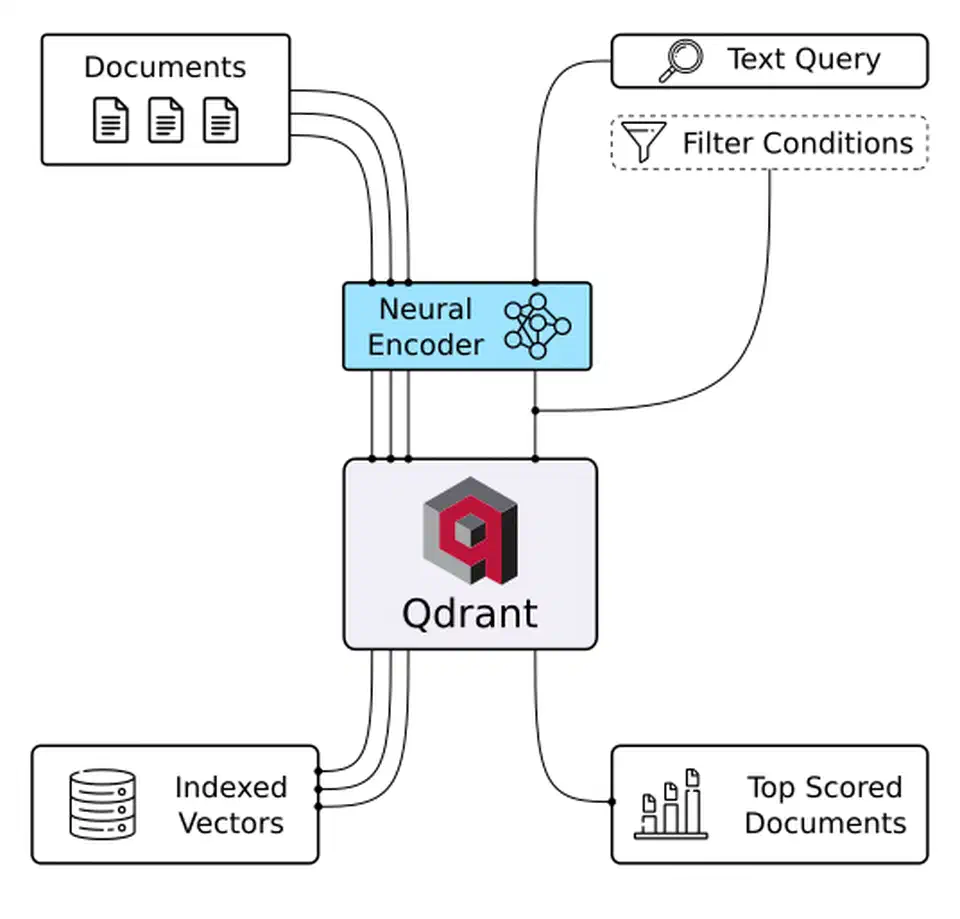
Hybrid search combines both methods with Reciprocal Rank Fusion (RRF). RRF merges ranked lists instead of raw scores, which avoids mismatched scales. It is simple, robust, and often beats either method alone on retrieval benchmarks. Qdrant supports hybrid search through sparse vectors. You can pair it with BGE-M3’s sparse output:
from qdrant_client.models import (
SparseVector,
NamedSparseVector,
NamedVector,
SearchRequest,
)
def hybrid_search(query: str, top_k: int = 10) -> list:
# Get both dense and sparse embeddings from BGE-M3
dense_embedding = list(embedder.embed([query]))[0].tolist()
# For sparse embeddings, use fastembed's sparse model
from fastembed import SparseTextEmbedding
sparse_embedder = SparseTextEmbedding(
model_name="Qdrant/bm42-all-minilm-l6-v2-attentions"
)
sparse_result = list(sparse_embedder.embed([query]))[0]
results = client.search_batch(
collection_name=COLLECTION_NAME,
requests=[
SearchRequest(
vector=NamedVector(name="dense", vector=dense_embedding),
limit=top_k,
with_payload=True,
),
SearchRequest(
vector=NamedSparseVector(
name="sparse",
vector=SparseVector(
indices=sparse_result.indices.tolist(),
values=sparse_result.values.tolist(),
),
),
limit=top_k,
with_payload=True,
),
],
)
# Apply RRF manually if Qdrant's built-in fusion isn't configured
scores = {}
for rank, hit in enumerate(results[0]):
scores[hit.id] = scores.get(hit.id, 0) + 1.0 / (60 + rank + 1)
for rank, hit in enumerate(results[1]):
scores[hit.id] = scores.get(hit.id, 0) + 1.0 / (60 + rank + 1)
# Sort by combined RRF score and retrieve payloads
sorted_ids = sorted(scores, key=scores.get, reverse=True)[:top_k]
return sorted_idsReranking gives you a second pass that improves precision. After hybrid search returns the top 20 candidates, a cross encoder scores each query and passage pair more accurately. A model like cross-encoder/ms-marco-MiniLM-L-6-v2 then trims the list to the top 4 results. That usually improves answer quality.
from sentence_transformers import CrossEncoder
reranker = CrossEncoder("cross-encoder/ms-marco-MiniLM-L-6-v2")
def rerank(query: str, candidates: list[dict], top_n: int = 4) -> list[dict]:
pairs = [(query, c["text"]) for c in candidates]
scores = reranker.predict(pairs)
ranked = sorted(zip(scores, candidates), key=lambda x: x[0], reverse=True)
return [doc for _, doc in ranked[:top_n]]Querying: Putting It All Together With Ollama
Once ingestion and retrieval work, the query pipeline is simple. Pull the top reranked chunks, format them as context, and send them to your local Ollama model with a prompt that tells it to answer only from the supplied material:
import ollama
def query_knowledge_base(question: str, model: str = "mistral-nemo") -> str:
# 1. Embed the question
question_embedding = list(embedder.embed([question]))[0].tolist()
# 2. Retrieve top candidates from Qdrant
search_results = client.search(
collection_name=COLLECTION_NAME,
query_vector=question_embedding,
limit=20,
with_payload=True,
)
candidates = [
{
"text": r.payload["text"],
"source": r.payload["source"],
"page": r.payload["page"],
}
for r in search_results
]
# 3. Rerank to top 4
top_chunks = rerank(question, candidates, top_n=4)
# 4. Build grounded context
context_parts = []
for i, chunk in enumerate(top_chunks, start=1):
context_parts.append(
f"[{i}] Source: {chunk['source']}, Page {chunk['page']}\n{chunk['text']}"
)
context = "\n\n---\n\n".join(context_parts)
# 5. Generate answer with Ollama
prompt = f"""You are a helpful assistant. Answer the user's question based ONLY on the provided context.
If the context does not contain sufficient information to answer, say so explicitly.
Cite the source number (e.g., [1], [2]) when referencing specific information.
Context:
{context}
Question: {question}
Answer:"""
response = ollama.chat(
model=model,
messages=[{"role": "user", "content": prompt}],
)
return response["message"]["content"]
# Example usage
answer = query_knowledge_base(
"What are the termination clauses in the vendor contract?"
)
print(answer)This is the complete loop: document in, answer out, entirely on your own hardware.
Local Model Context Protocol (MCP) Servers
Running the query pipeline as a one off Python script works for one person. It gets limiting when you want the same knowledge base in several tools, such as an Obsidian plugin, a Claude Desktop session, Open WebUI, or a custom CLI. The Model Context Protocol (MCP) fixes that by turning the RAG pipeline into a small service that any MCP client can call as a tool.
An MCP server listens for structured tool calls and returns structured responses. To an LLM client, calling your RAG knowledge base looks like any other MCP tool. The client states what it needs. MCP routes the call. Your server runs the Qdrant query and returns the retrieved chunks. The LLM does not need to know that vector search happened underneath.
Setting up a minimal MCP server with the mcp Python library takes about 50 lines of code:
from mcp.server import Server
from mcp.server.stdio import stdio_server
from mcp import types
import asyncio
app = Server("rag-knowledge-base")
@app.list_tools()
async def list_tools():
return [
types.Tool(
name="search_knowledge_base",
description="Search the private local knowledge base for information relevant to a query.",
inputSchema={
"type": "object",
"properties": {
"query": {
"type": "string",
"description": "The question or search query",
},
"top_k": {
"type": "integer",
"description": "Number of results to return (default 4)",
"default": 4,
},
},
"required": ["query"],
},
)
]
@app.call_tool()
async def call_tool(name: str, arguments: dict):
if name == "search_knowledge_base":
query = arguments["query"]
top_k = arguments.get("top_k", 4)
embedding = list(embedder.embed([query]))[0].tolist()
results = client.search(
collection_name=COLLECTION_NAME,
query_vector=embedding,
limit=top_k * 3,
with_payload=True,
)
candidates = [
{
"text": r.payload["text"],
"source": r.payload["source"],
"page": r.payload["page"],
}
for r in results
]
top_chunks = rerank(query, candidates, top_n=top_k)
output = "\n\n".join(
f"[Source: {c['source']}, p.{c['page']}]\n{c['text']}" for c in top_chunks
)
return [types.TextContent(type="text", text=output)]
async def main():
async with stdio_server() as streams:
await app.run(*streams, app.create_initialization_options())
if __name__ == "__main__":
asyncio.run(main())With this MCP server running, you can connect it to Claude Desktop by adding it to claude_desktop_config.json, or to Open WebUI’s tool settings. Your private Qdrant knowledge base then becomes a tool any LLM session can use without cloud services.
For Obsidian users, pairing this with obsidian-local-rest-api (a community plugin that exposes your vault’s notes as a REST API) lets you keep your vault indexed in Qdrant and query it from any MCP client. The choice is simple. Build an MCP server when you want to reuse the RAG system across many tools and clients. Skip it when you only need one purpose built script.
Privacy and Zero-Cloud Architecture
The entire point of a local stack is privacy. Still, “it runs locally” is not a full privacy plan. You need to think about a few clear threat paths.
One common privacy failure is using a cloud LLM for generation. You might store private HR files in local Qdrant and retrieve chunks on your machine. If you then send those chunks to the OpenAI API, the data has still left your machine. Keep the final answer step local with Ollama, even during development.
The second threat is accidental PII in your documents. If files contain employee names, email addresses, patient records, or financial account numbers, those strings can land in the vector store and later appear in retrieved chunks. The presidio library from Microsoft gives you a fast local pipeline for PII detection and redaction. Run it before chunking:
from presidio_analyzer import AnalyzerEngine
from presidio_anonymizer import AnonymizerEngine
analyzer = AnalyzerEngine()
anonymizer = AnonymizerEngine()
def redact_pii(text: str) -> str:
results = analyzer.analyze(text=text, language="en")
anonymized = anonymizer.anonymize(text=text, analyzer_results=results)
return anonymized.text
# Use in your ingestion pipeline before chunking
clean_text = redact_pii(raw_document_text)
chunks = splitter.split_text(clean_text)Network isolation helps block accidental outbound calls from Docker containers. Qdrant is local, but telemetry is on by default. Turn it off:
docker run -d --name qdrant \
-p 6333:6333 -p 6334:6334 \
-v $(pwd)/qdrant_storage:/qdrant/storage \
-e QDRANT__TELEMETRY_DISABLED=true \
qdrant/qdrantFor maximum isolation, run the whole RAG stack inside a Docker Compose network with no external access. Add firewall rules so the embedding and LLM processes cannot make outbound connections. A minimal docker-compose.yml with network isolation:
version: "3.9"
services:
qdrant:
image: qdrant/qdrant
ports:
- "6333:6333"
volumes:
- ./qdrant_storage:/qdrant/storage
environment:
- QDRANT__TELEMETRY_DISABLED=true
networks:
- rag_internal
networks:
rag_internal:
driver: bridge
internal: true # no external internet accessHardware affects the experience more than many guides admit. Qdrant loads its HNSW index from disk on startup. On a 5400 RPM HDD, a large index can take tens of seconds to load, which hurts cold start latency. On a Gen4 NVMe SSD, the same load can finish in under a second. If you plan to store tens of thousands of chunks, put the Qdrant volume on NVMe. Once the index is warm, query latency usually falls to single-digit milliseconds.
Evaluating and Maintaining Your Knowledge Base
If you cannot measure your RAG system, you cannot improve it. The ragas library gives you metrics built for RAG without human labels: context precision, answer faithfulness, and answer relevancy. Running even 20-50 test question and answer pairs through ragas will show whether your chunking, retrieval settings, or reranker thresholds need work.
Adding new documents is simple: ingest the new files and upsert the new points into the current collection. Qdrant upsert is idempotent when you use deterministic IDs, such as a hash of the source path and chunk index. To handle deletions, keep a small SQLite table that maps (source_path, point_id).
Re-embedding changed files is the tricky part. If you update a document and ingest it again without deleting the old chunks, you end up with both versions in the index. Delete all points for that source file first, then upsert the new chunks. If you stored the source path in the payload, Qdrant payload filters make this one delete call:
from qdrant_client.models import Filter, FieldCondition, MatchValue
client.delete(
collection_name=COLLECTION_NAME,
points_selector=Filter(
must=[
FieldCondition(key="source", match=MatchValue(value="updated_report.pdf"))
]
),
)
# Then re-ingest the updated file
ingest_pdf(Path("./documents/updated_report.pdf"))The result is a knowledge base that stays current with your documents, runs on your own hardware, and gives your local LLM grounded context instead of forcing it to guess. The full stack, Qdrant, BGE-M3, and Ollama, needs no cloud accounts, API keys, or monthly fees. Your documents stay where you put them.
 Botmonster Tech
Botmonster Tech